goat anti gal3 (R&D Systems)
Structured Review

Goat Anti Gal3, supplied by R&D Systems, used in various techniques. Bioz Stars score: 93/100, based on 95 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more
https://www.bioz.com/result/goat anti gal3/product/R&D Systems
Average 93 stars, based on 95 article reviews
Images
1) Product Images from "Microglia at Sites of Atrophy Restrict the Progression of Retinal Degeneration via Galectin-3 and Trem2 Interactions"
Article Title: Microglia at Sites of Atrophy Restrict the Progression of Retinal Degeneration via Galectin-3 and Trem2 Interactions
Journal: bioRxiv
doi: 10.1101/2023.07.19.549403

Figure Legend Snippet: ( A ) UMAP plot showing integrated clustering of immune cells samples from four mouse models of retinal degeneration, including LD model (sorted by Cx3cr1 + ), NaIO 3 model (CD45 + ), P23H model (CD45 + ) and aging model (CD45 + ) and naïve mice (CD45 + ). A total of 15,623 macrophages, including 13,489 microglia, were integrated among four models. PMN, polymorphonuclear neutrophils; mo-MFs, monocyte-derived macrophages; pv-MFs: perivascular macrophages; NK, natural killer. ( B ) UMAP plots showing integrated macrophage clusters by two datasets. Dash circles indicate subretinal microglia (srMG). ( C ) Percentage of sample distribution by clusters. The arrow indicates the enrichment of srMG cluster from degenerating retinas. ( D ) Heatmap of top 30 conserved marker genes of subretinal microglia shared by each model across clusters. Genes were ranked by fold changes. Arrows indicate srMG cluster. ( E ) In situ validation of Gal3 expression on the apical RPE (top) or in the neuroretina from the inner plexiform layer (bottom). Iba1 (green), phalloidin (red, only in RPE) and Gal3 (magenta). Scale bar: 100μm. ( F ) Percentage of Gal3 + cells relative to Iba1 + cells between RPE and neuroretina tissues across models.
Techniques Used: Derivative Assay, Marker, In Situ, Expressing

Figure Legend Snippet: ( A ) Iba1 (green) and phalloidin (red) staining in RPE flatmounts from LD-subjected mice as indicated. ( B ) Quantifications of subretinal Iba1 + cells as shown in A. ( C ) Iba1 (green) and phalloidin (red) staining in RPE flatmounts from P23H mice as indicated. ( D ) Quantifications of subretinal Iba1 + cells as shown in C. ( E ) Examples of ERG responses at different flash intensities as indicated. ( F ) Representative retinal cross sections of WT, Lgal3 +/- and Lgal3 -/- in P23H mice. ( G and H ) Quantifications of Gal3 depletion efficiency (G) and frequencies of subretinal Iba1 + cells (H) in Gal3 cKO mice (n=9) compared with genotype control mice (n=9) and tamoxifen control (n=8). Scale bars: 100 μm. Data were collected from 2-3 independent experiments. ***: p<0.001; ns: not significant (one-way ANOVA with Tukey’s post hoc test).
Techniques Used: Staining

Figure Legend Snippet: ( A ) Images of phalloidin staining in WT and Lgal3 -/- RPE tissues in LD. ( B ) Quantifications of dysmorphic RPE cells (n=6, 7 and 3, respectively). ( C ) TUNEL (green) and DAPI (blue) staining in WT and Lgal3 -/- retinal cross sections in LD. ONL and INL, outer and inner nuclear layers. ( D ) Quantifications of TUNEL + photoreceptors in ONL (n=5, 5 and 3, respectively). ( E ) Rhodopsin (red) and Iba1 (green) staining in WT and Lgal3 -/- retinal cross sections in LD. Images from single planes of confocal scans were shown. ( F ) Quantifications of rhodopsin+ subretinal microglia (n=4 per group). ( G ) Images of phalloidin staining in WT and Lgal3 -/- RPE tissues at 2 years of age. ( H ) Quantifications of RPE cell size. Dots represent individual images with n=5 mice per group. ( I ) ERG data showing scotopic a- and b-waves in 2-year-old WT (n=5) and Lgal3 -/- (n=5) mice. ( J ) Scotopic a- and b-waves of ERG data among Lgal3 +/+ (n=12), Lgal3 +/- (n=6) and Lgal3 -/- (n=10) in P23H mice. ( K ) Quantifications of ONL thickness among Lgal3 +/+ (n=7), Lgal3 +/- (n=7), and Lgal3 -/- (n=8) in P23H mice. ( L ) Representative images of dysmorphic RPE cells in Gal3 cKO in LD. Iba1, green; phalloidin, red; Gal3, magenta. ( M ) Quantifications of dysmorphic RPE cells in Gal3 cKO mice (n=9) compared with genotype control ( Cx3cr1 CreER/+ Lgals3 fl/fl mice, n=9) and tamoxifen control ( Cx3cr1 CreER/+ mice treated with tamoxifen, n=8). Scale bars: 100μm. Data were collected from 2-3 independent experiments. *: p<0.05; **: p<0.01; ***: p<0.001. One-way ANOVA with Tukey’s post hoc test (B, D and M); unpaired Student’s t-test (F and H); two-way ANOVA with Tukey’s post hoc test (I, J and K).
Techniques Used: Staining, TUNEL Assay
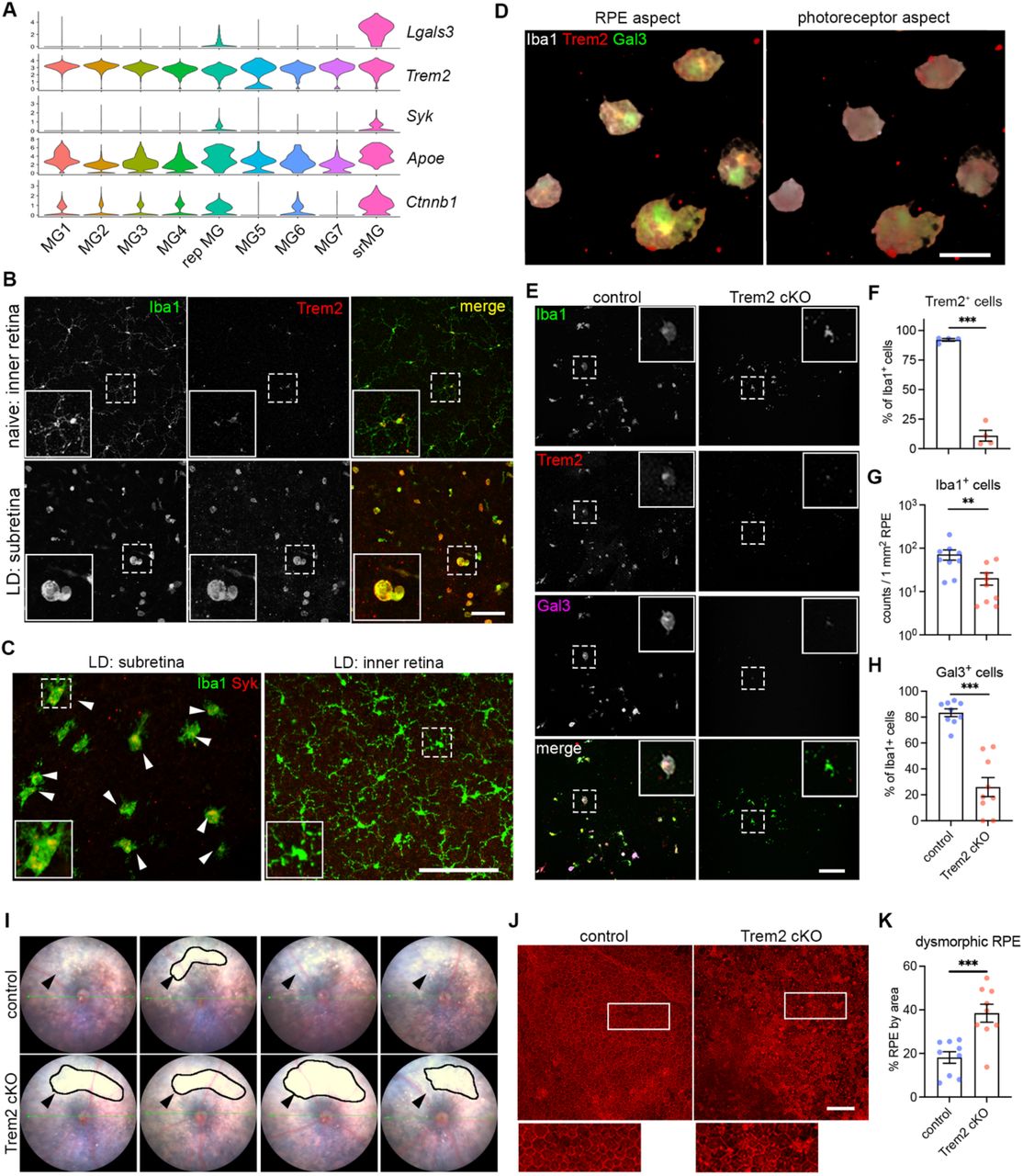
Figure Legend Snippet: ( A ) Violin plots showing the upregulation of genes ( Lgals3, Syk and Ctnnb1I) related to Trem2 signaling by subretinal microglia from the integrated dataset of all four mouse models. ( B ) Images of Iba1 (green) and Trem2 (red) staining in naïve microglia from inner retina and subretinal microglia in LD. ( C ) Images of Iba1 (green) and Syk (red) staining in subretinal microglia and microglia from inner retina in LD. ( D ) 3D rendering images of Gal3 (green), Trem2 (red) and Iba1 (white) staining in subretinal microglia in LD. Views from both the apical RPE aspect and neuroretina aspect are shown. ( E ) Images of Iba1 (green), Trem2 (red) and Gal3 (magenta) staining in subretinal microglia between control and Trem2 cKO mice in LD. ( F-H ) Quantifications of Trem2 depletion (F, n=4 per group), Iba1 + cells (G, n=9) and Gal3 + cells (H, n=9) between control and Trem2 cKO mice. ( I ) Fundus images showing increased subretinal white lesions in of Trem2 cKO mice in LD as indicated by arrows. Images from four individual mice per group are shown. ( J ) Images of phalloidin staining in RPE tissues from control and Trem2 cKO mice in LD. ( K ) Quantifications of dysmorphic RPE cells between control and Trem2 cKO mice (n=9 per group). Scale bars: 50μm (D); 100μm (B, C E,and J). Data were collected from 2 independent experiments. **: p<0.01; ***: p<0.001. Unpaired Student’s t-test (F-H).
Techniques Used: Staining

Figure Legend Snippet: ( A ) Split views of confocal scans showing the colocalization of Trem2 (red) and Gal3 (green) in the subretinal microglia. Lines indicate the RPE-facing and neuroretina (NR)-facing aspects as indicated. ( B ) Fundus images showing increased subretinal white lesions in anti-Trem2 mAb178 treated mice in LD as indicated by arrows. Images of 4 individual mice per group are shown. ( C ) Images of Iba1 (green) and Gal3 (magenta) staining in subretinal microglia between control and mAb178-treated mice in LD. Scale bar: 100 μm. ( D and E ) Quantifications of Iba1 + cells and Gal3 + cells between control and mAb178 (n=8 per group). ( F ) Images of phalloidin staining in RPE flatmounts from control and mAb178 treated mice in LD. Scale bar: 100μm. ( G ) Quantifications of dysmorphic RPE cells between control (n=8) and mAb178 (n=9) treated mice. ( H ) Images of Iba1 (green) and Trem2 (red) in microglia from the inner retina of naïve control and Trem2 cKO mice. Scale bar: 50μm.
Techniques Used: Staining

Figure Legend Snippet: ( A ) ELISA of soluble Trem2 (sTrem2) in vitreous fluid and retinal fluid from naïve WT mice, WT and Trem2 cKO mice subjected to LD. ( B ) Fundus images of mice treated with isotype control or 4D9 anti-Trem2 in LD. Four individual mice per group are shown. ( C ) Representative OCT images of mice treated with isotype or 4D9 in LD. ( D ) Quantifications of outer nuclear layer (ONL) thickness by OCT (n=13 per group). ONL thickness was measured at both nasal and temporal sides. ( E and F ) Scotopic a-waves and b-waves of ERG data among mice treated with isotype or 4D9 in naïve or LD setting (n=5 per group). ( G ) Fundus images of Gal3 cKO mice treated with isotype or 4D9 in LD. Four individual mice per group are shown. ( H ) Representative OCT images of Gal3 cKO mice treated with isotype control or 4D9 anti-Trem2 in LD. ( I ) Quantifications of average ONL thickness by OCT between control and Gal3 cKO mice treated with either isotype or 4D9 (n=13 per group). ( J ) Images of phalloidin staining of control and Gal3 cKO RPE treated with isotype or 4D9 in LD. ( K ) Quantifications of dysmorphic RPE cells (n=15, 13, 11 and 13, respectively). Scale bars: 100μm. Data were collected from 2-4 independent experiments. *: p<0.05; **: p<0.01; ***: p<0.001. Unpaired Student’s t-test (F-H). One-way ANOVA with Tukey’s post hoc test (A); two-way ANOVA with Tukey’s post hoc test (D-F, I and K).
Techniques Used: Enzyme-linked Immunosorbent Assay, Staining

Figure Legend Snippet: ( A ) Staining of human IgG (red) and Iba1 (green) in retinal cross sections collected from mice with or without 4D9 treatment in LD. The hIgG is used to trace 4D9 antibodies, which outlines retinal vasculatures in 4D9 treated mice. Arrows indicate the presence of 4D9 antibodies in the subretinal microglia, while asters indicate the absence of 4D9 antibodies in microglia from the inner retina. ( B ) Human IgG (red) and Iba1 (green) staining in RPE and neuroretina flatmounts collected from mice treated with 4D9 antibodies in LD. ( C ) Quantifications of hIgG + microglia in the subretinal space and neuroretina. ( D and E ) Quantifications of Iba1 + cells and Gal3 + cells between control and Gal3 cKO mice treated with either isotype or 4D9 (n=13 per group). Scale bars: 100 μm. Data were collected from 2-4 independent experiments. ***: p<0.001; ns: not significant (unpaired Student’s t-test: C; two-way ANOVA with Tukey’s post hoc test: D and E).
Techniques Used: Staining

Figure Legend Snippet: ( A ) Multispectral imaging of GAL3 and CD68 co-staining in the subretinal space (top) and inner retina (bottom) from human donors. Unmixed purple spectrum (GAL3) and yellow spectrum (CD68) are shown. The areas of colocalized spectra are highlighted in green. Scale bar: 50μm. ONL and INL, outer and inner nuclear layers. ( B ) Representative image of Gal3 and CD68 co-staining in the macular GA region of a retinal section from an 88-year-old female donor eye with advanced AMD (Sarks V). Black insert box shows the magnification of GA with double positive cells. Scale bar: 200μm. ONL and INL, outer and inner nuclear layers; GCL, ganglion cell layer. ( C ) Correlation between the frequencies of macular Gal3 + CD68 + double positive cells (y axis) and Sarks AMD grading (x axis) by Spearman’s correlation (n = 18 donors, Table S2). Coefficient and p-value are shown. ( D ) Histograms showing increased TREM2 + myeloid cells (CD45 + CD11B + ) in RPE/choroid tissues of AMD donors. Concatenated histograms were shown (n=3 per groups). Control human blood samples were used to set up flow gating. ( E-G ) Quantifications of TREM2 + (E), CD45 + (F), and CD11B + (G) cell frequencies in RPE/choroid tissues between non-AMD and AMD donors. Unpaired Student’s t test is used. P-values are shown. ( H ) Correlation between the frequencies of TREM2 + myeloid cells (y axis) and Sarks AMD grading (x axis) in RPE/choroid tissues by Spearman’s correlation. Coefficient and p-value are shown.
Techniques Used: Imaging, Staining

Figure Legend Snippet: ( A ) Images of GAL3 (purple) and CD68 (yellow) co-staining in the macula region of retinal sections from human donors categorized by Sark grades (I-VI). The macular neurosensory retinas of some subject eyes exhibited fixation-related artifactual detachment. In these subjects, separate images of RPE/choroid tissues are shown. Scale bar: 100μm. ONL and INL, outer and inner nuclear layers. GCL, ganglion cell layer. ( B ) Spectral imaging of GAL3 and CD68 co-staining in the geographic atrophy from donor #23 with advanced AMD (Sarks V). Unmixed purple spectrum (GAL3) and yellow spectrum (CD68) are shown. The areas of colocalized spectra are highlighted in green. Scale bar: 50μm. ( C and D ) Images showing the presence of subretinal GAL3 (purple) and CD68 (yellow) double positive cells in the areas with photoreceptor loss and preserved RPE in the transitional area of the macula from an AMD donor (C) and in the age-related peripheral degeneration of a non-AMD donor (D). Scale bars: 100μm. ( E ) Gating strategy of flow cytometry analysis. CD45 + CD11B + cells and CD45 + CD11B - cells from control blood were used to determine the gating of TREM2 + cells. Concatenated plots are shown for non-AMD and AMD. ( F ) Flow contour plots of individual donors showing increased percentage of TREM2 + myeloid cells in AMD.
Techniques Used: Staining, Imaging, Flow Cytometry
























